(702) Evaluation of LABType® CWD Versus XR Beads for HLA Typing: Implications for Transition in Clinical Workflow
- JL
Juan Liu, PhD
– Assistant Professor, University of Arkansas for Medical Sciences
Speaker(s)
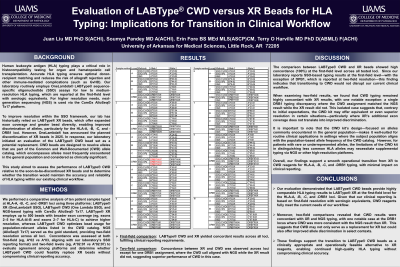
Aim: Our laboratory routinely performs low to medium resolution HLA typing using One Lambda’s LABType® sequence-specific oligonucleotide (SSO) assay, supplemented by high-resolution typing via next-generation sequencing (NGS) using CareDx®’s AlloSeq® Tx 17 platform. For HLA-A, -B, -C, and -DRB1 loci, LABType XR beads have been employed to achieve higher resolution within the SSO framework. However, One Lambda has announced the planned discontinuation of XR beads in 2025, prompting the need to evaluate the performance of LABType CWD beads as a replacement.
Methods: To evaluate assay comparability, we performed HLA typing on 10 patient samples at the HLA-A, -B, -C, and -DRB1 loci using three platforms: LABType® XR, LABType® CWD, and NGS. According to One Lambda®, the XR kit provides enhanced exon coverage (eg, exons 2-5 for A and B loci and exons 2-7 for C locus) and higher resolution by utilizing up to 500 bead regions, while the CWD kit resolves alleles based on the Common and Well-Documented (CWD) catalog.
Results: In our current clinical workflow, although NGS results are reported at the two-field level, LABType® SSO typing is reported at the first-field level with serologic equivalents - except for DPB1, which is reported at the two-field level. Based on this reporting policy, our data show that LABType® CWD and XR yield comparable results at the first-field level for HLA-A, -B, -C, and -DRB1 loci, meeting our laboratory’s clinical reporting requirements. Notably, when evaluated at the two-field level, results were concordant between XR and CWD across all loci except for one DRB1 assignment, where the CWD result matched the NGS call, but the XR result did not. This observation suggests that, at least in this instance, the CWD assay may outperform XR in DRB1 resolution.
Conclusion: These findings support a smooth transition to LABType® CWD typing reagents for HLA-A, -B, -C, and -DRB1 loci in clinical practice, particularly when reporting is limited to the first-field and serologic equivalent. Further evaluation across a larger sample set may help confirm performance at higher resolution and across a broader allele spectrum.
Methods: To evaluate assay comparability, we performed HLA typing on 10 patient samples at the HLA-A, -B, -C, and -DRB1 loci using three platforms: LABType® XR, LABType® CWD, and NGS. According to One Lambda®, the XR kit provides enhanced exon coverage (eg, exons 2-5 for A and B loci and exons 2-7 for C locus) and higher resolution by utilizing up to 500 bead regions, while the CWD kit resolves alleles based on the Common and Well-Documented (CWD) catalog.
Results: In our current clinical workflow, although NGS results are reported at the two-field level, LABType® SSO typing is reported at the first-field level with serologic equivalents - except for DPB1, which is reported at the two-field level. Based on this reporting policy, our data show that LABType® CWD and XR yield comparable results at the first-field level for HLA-A, -B, -C, and -DRB1 loci, meeting our laboratory’s clinical reporting requirements. Notably, when evaluated at the two-field level, results were concordant between XR and CWD across all loci except for one DRB1 assignment, where the CWD result matched the NGS call, but the XR result did not. This observation suggests that, at least in this instance, the CWD assay may outperform XR in DRB1 resolution.
Conclusion: These findings support a smooth transition to LABType® CWD typing reagents for HLA-A, -B, -C, and -DRB1 loci in clinical practice, particularly when reporting is limited to the first-field and serologic equivalent. Further evaluation across a larger sample set may help confirm performance at higher resolution and across a broader allele spectrum.
