(601) KIR Complete MX: Multiplexed whole gene genotyping of all KIR genes using ONT sequencing

Maarten Penning, PhD
– General Manager, GenDx
Speaker(s)
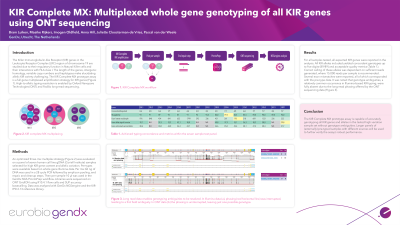
Aim: The Killer Immunoglobulin-like Receptor (KIR) genes located in the Leukocyte Receptor Complex (LRC) region of chromosome 19 have been subject to extensive study because of their regulatory function in Natural Killer cells and their interactions with class I HLA. Due to the length of the genes, high intergenic homology and the highly variable haplotypes, elucidating allelic KIR variety remains challenging. To create a full gene amplification strategy that allows for high-to-allelic resolution KIR typing, we developed KIR Complete MX. This prototype strategy is compatible with long-read sequencing on Oxford Nanopore Technologies (ONT) sequencers.
Methods: We developed an amplification strategy for all KIR genes, including pseudogenes KIR2DP1 and -3DP1. To balance set-up efficiency with minimal co- and cross-amplifications, a 3 tube multiplex strategy gave the best results. To evaluate the performance, a panel of 7 samples, pre-typed using singleplex whole gene typing on Illumina sequencing, was evaluated. The panel was selected for high gene content and allelic variation. Amplification by KIR Complete MX was followed by end preparation and cleanup steps before samples were used in the GenDx NGS-ProntoPrep workflow. The generated libraries were sequenced using the ONT GridION platform and resulting data was analyzed with GenDx NGSengine software and KIR-IPD 2.13 reference library.
Results: For all samples tested, all expected KIR genes were reported in the analysis. Correct calling of loci was dependent on ample read generation. Several loci required additional investigation and the introduction of optimized NGSengine settings for the analysis, to yield the correct alleles. With these adjustments, all alleles were in concordance with the pre-types. It was noted that genotype ambiguities, a common occurrence in Illumina based KIR typing, were fully absent due to the long-read phasing possibilities offered by the ONT sequencing data.
Conclusion: The presented method is capable of accurately genotyping all KIR genes and alleles in the tested high variation sample set without genotype ambiguities. Expansion of these initial tests to larger sample panels is required to further validate this assay.
Methods: We developed an amplification strategy for all KIR genes, including pseudogenes KIR2DP1 and -3DP1. To balance set-up efficiency with minimal co- and cross-amplifications, a 3 tube multiplex strategy gave the best results. To evaluate the performance, a panel of 7 samples, pre-typed using singleplex whole gene typing on Illumina sequencing, was evaluated. The panel was selected for high gene content and allelic variation. Amplification by KIR Complete MX was followed by end preparation and cleanup steps before samples were used in the GenDx NGS-ProntoPrep workflow. The generated libraries were sequenced using the ONT GridION platform and resulting data was analyzed with GenDx NGSengine software and KIR-IPD 2.13 reference library.
Results: For all samples tested, all expected KIR genes were reported in the analysis. Correct calling of loci was dependent on ample read generation. Several loci required additional investigation and the introduction of optimized NGSengine settings for the analysis, to yield the correct alleles. With these adjustments, all alleles were in concordance with the pre-types. It was noted that genotype ambiguities, a common occurrence in Illumina based KIR typing, were fully absent due to the long-read phasing possibilities offered by the ONT sequencing data.
Conclusion: The presented method is capable of accurately genotyping all KIR genes and alleles in the tested high variation sample set without genotype ambiguities. Expansion of these initial tests to larger sample panels is required to further validate this assay.
