(315) Assessment of Chimerism Using Two Next Generation Sequencing Based Methods

Jasmine Kendrick, MPH, MLS(ASCP), CHT(ACHI),
– Development Technologist, Mayo Clinic
Speaker(s)
Aim: Short tandem repeat (STR) based chimerism testing has been the gold standard for assessment of engraftment after allogeneic hematopoietic stem cell transplant (allo HSCT). In this study, two next generation sequencing (NGS) based multiplex methods using either insertions/deletions (indels) or single nucleotide polymorphisms (SNPs) were compared.
Methods: DNA extracted from peripheral blood of 10 healthy volunteers were used to create a broad range of pre-defined mixed chimerism percentages and establish a reportable range. Extracted DNA from 19 post-HCT samples were tested by both NGS-based chimerism methods and compared to the reportable range to assess concordance.
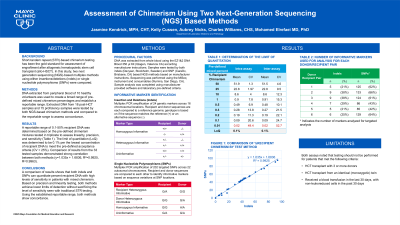
Results: A reportable range of 0.1-50% recipient DNA was determined based on the pre-defined chimerism mixtures tested in triplicate to assess linearity, precision, and sensitivity (Table 1). The limit of quantitation (LoQ) was determined to be 0.1% per the lowest concentration of recipient DNA to meet the pre-defined acceptance criteria (CV < 25%). Comparison of results from the 19 post-HCT samples (table 2) demonstrated strong correlation between both methods (y=0.9494x + 2.669, R2=0.9358, R=0.9674, p=0.33).
Conclusion: A comparison of results shows that both indels and SNPs can quantitate percent recipient DNA with high levels of sensitivity in patients with mixed chimerism. Based on precision and linearity testing, both methods achieve lower limits of detection without sacrificing the level of sensitivity seen with traditional STR testing. Using the established reportable range, both methods show concordance.
Methods: DNA extracted from peripheral blood of 10 healthy volunteers were used to create a broad range of pre-defined mixed chimerism percentages and establish a reportable range. Extracted DNA from 19 post-HCT samples were tested by both NGS-based chimerism methods and compared to the reportable range to assess concordance.
Results: A reportable range of 0.1-50% recipient DNA was determined based on the pre-defined chimerism mixtures tested in triplicate to assess linearity, precision, and sensitivity (Table 1). The limit of quantitation (LoQ) was determined to be 0.1% per the lowest concentration of recipient DNA to meet the pre-defined acceptance criteria (CV < 25%). Comparison of results from the 19 post-HCT samples (table 2) demonstrated strong correlation between both methods (y=0.9494x + 2.669, R2=0.9358, R=0.9674, p=0.33).
Conclusion: A comparison of results shows that both indels and SNPs can quantitate percent recipient DNA with high levels of sensitivity in patients with mixed chimerism. Based on precision and linearity testing, both methods achieve lower limits of detection without sacrificing the level of sensitivity seen with traditional STR testing. Using the established reportable range, both methods show concordance.
