(211) HLA Molecular Mismatch Categories Revised and Validated Using Updated HLAMatchmaker Software

Emmett Wong, MBBS, MMed (Int Med), MRCP (UK), FRCP (Edin), MCI, FAMS (Renal Medicine), FASN
– Adjunct Assistant Professor, National University of Singapore, Singapore
Speaker(s)
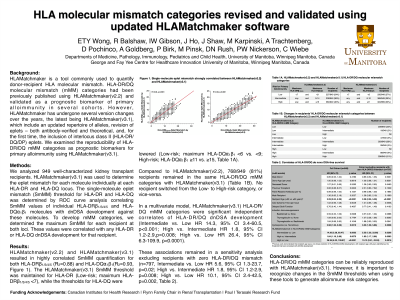
Aim: HLAMatchmaker is commonly used to quantify donor-recipient HLA molecular mismatch. HLA-DR/DQ molecular mismatch (mMM) categories had been previously published using HLAMatchmaker(v2.2) and validated as a prognostic biomarker of primary alloimmunity in several cohorts. However, HLAMatchmaker has undergone several version changes over the years, the latest being HLAMatchmaker(v3.1). We examined the reproducibility of HLA-DR/DQ mMM categories as prognostic biomarkers for primary alloimmunity using HLAMatchmaker(v3.1).
Methods: We analyzed 949 well-characterized kidney transplant recipients. HLAMatchmaker(v3.1) was used to determine the eplet mismatch for each molecule individually at each HLA-DR and HLA-DQ locus. The single-molecule eplet mismatch (SmMM) threshold for HLA-DR and HLA-DQ was determined by ROC curve analysis correlating SmMM values of individual HLA-DRb1/3/4/5 and HLA-DQa1b1 molecules with dnDSA development against these molecules. To develop mMM categories, we determined the maximum SmMM for each recipient at both loci. These values were correlated with any HLA-DR or HLA-DQ dnDSA development for that recipient.
Results: HLAMatchmaker(v2.2) and HLAMatchmaker(v3.1) resulted in highly correlated SmMM quantification for both HLA-DRβ1/3/4/5 (R2=0.88) and HLA-DQa1b1 (R2=0.93). The HLAMatchmaker(v3.1) SmMM threshold was maintained for HLA-DR (Low-risk; maximum HLA-DRβ1/3/4/5 <7), while the thresholds for HLA-DQ were lowered (Low-risk; maximum HLA-DQa1b1 <6 vs. <9; High-risk; HLA-DQa1b1 ≥11 vs. ≥15, Table 1A). Compared to HLAMatchmaker(v2.2), 768/949 (81%) recipients remained in the same HLA-DR/DQ mMM categories with HLAMatchmaker(v3.1) (Table 1B). No recipient switched from the Low- to High-risk category, or vice-versa. In a multivariate model, HLAMatchmaker(v3.1) HLA-DR/DQ mMM categories were significant independent correlates of HLA-DR/DQ dnDSA development (Intermediate vs. Low HR 14.3, 95% CI 3.4-60.5, p<0.001; High vs. Intermediate HR 1.8, 95% CI 1.2-2.9, p=0.008; High vs. Low HR 26.4, 95% CI 6.3-109.9, p<0.0001). These associations remained in a sensitivity analysis excluding recipients with zero HLA-DR/DQ mismatch (n=797, Table 2).
Conclusion: HLA-DR/DQ mMM categories can be reliably reproduced with HLAMatchmaker(v3.1). However, it is important to recognize changes in the SmMM thresholds when using these tools to generate alloimmune risk categories.
Methods: We analyzed 949 well-characterized kidney transplant recipients. HLAMatchmaker(v3.1) was used to determine the eplet mismatch for each molecule individually at each HLA-DR and HLA-DQ locus. The single-molecule eplet mismatch (SmMM) threshold for HLA-DR and HLA-DQ was determined by ROC curve analysis correlating SmMM values of individual HLA-DRb1/3/4/5 and HLA-DQa1b1 molecules with dnDSA development against these molecules. To develop mMM categories, we determined the maximum SmMM for each recipient at both loci. These values were correlated with any HLA-DR or HLA-DQ dnDSA development for that recipient.
Results: HLAMatchmaker(v2.2) and HLAMatchmaker(v3.1) resulted in highly correlated SmMM quantification for both HLA-DRβ1/3/4/5 (R2=0.88) and HLA-DQa1b1 (R2=0.93). The HLAMatchmaker(v3.1) SmMM threshold was maintained for HLA-DR (Low-risk; maximum HLA-DRβ1/3/4/5 <7), while the thresholds for HLA-DQ were lowered (Low-risk; maximum HLA-DQa1b1 <6 vs. <9; High-risk; HLA-DQa1b1 ≥11 vs. ≥15, Table 1A). Compared to HLAMatchmaker(v2.2), 768/949 (81%) recipients remained in the same HLA-DR/DQ mMM categories with HLAMatchmaker(v3.1) (Table 1B). No recipient switched from the Low- to High-risk category, or vice-versa. In a multivariate model, HLAMatchmaker(v3.1) HLA-DR/DQ mMM categories were significant independent correlates of HLA-DR/DQ dnDSA development (Intermediate vs. Low HR 14.3, 95% CI 3.4-60.5, p<0.001; High vs. Intermediate HR 1.8, 95% CI 1.2-2.9, p=0.008; High vs. Low HR 26.4, 95% CI 6.3-109.9, p<0.0001). These associations remained in a sensitivity analysis excluding recipients with zero HLA-DR/DQ mismatch (n=797, Table 2).
Conclusion: HLA-DR/DQ mMM categories can be reliably reproduced with HLAMatchmaker(v3.1). However, it is important to recognize changes in the SmMM thresholds when using these tools to generate alloimmune risk categories.
