(200) HLA-DPB1 Allele Frequencies across Four Donor Populations
- NL
Nathan A. Lemp, PhD, F(ACHI)
– HLA Laboratory Director, Eurofins Donor & Product Testing, LLC, United States
Speaker(s)
Aim: The update to the OPTN HLA Equivalency tables in 12/2024 brought substantial changes to the HLA-DPB1 Unacceptable Antigen Equivalences, including the addition of p-groups and many rare alleles. We performed an analysis of the DPB1 alleles identified in four of our laboratories serving distinct regional donor populations across the United States to examine allele frequencies among these populations and determine the impact of the updated HLA Equivalence tables.
Methods: HLA typing was performed using LinkSeq HLA Typing Kits (One Lambda) as the primary method. To make rare (non-CWD 2.0) allele assignments, some specimens were additionally tested using Micro SSP (One Lambda), LABType SSO (One Lambda), and/or Olerup SSP (CareDx). 5,542 specimens tested between 01/2023 and 04/2025 at four Eurofins DPT laboratories (Honolulu (2,714), Los Angeles (1,218), Denver (502), and Philadelphia (1,108)) were included in the analysis (numbers in parentheses represent the volumes tested at each lab). Due to the lower numbers of deceased donors in Honolulu, living donor and transplant candidates were also included.
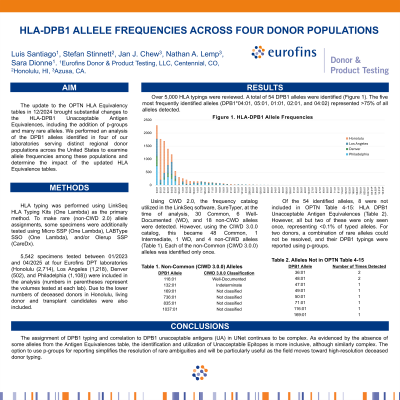
Results: Over 5,000 HLA typings were reviewed. A total of 54 DPB1 alleles were identified (Figure 1). The five most frequently identified alleles represented >75% of all alleles detected. Using CWD 2.0, the frequency catalog utilized in the LinkSeq software, SureTyper, 30 Common, 6 Well-Documented (WD), and 18 non-CWD alleles were detected. However, using the CIWD 3.0.0 catalog, this became 48 Common, 1 Intermediate, 1 WD, and 4 non-CIWD alleles. Each of the non-Common (CIWD 3.0.0) alleles was identified only once. Of the 54 identified alleles, 8 were not included in OPTN Table 4-15: HLA DPB1 Unacceptable Antigen Equivalences. However, all but two of these were only seen once, representing <0.1% of typed alleles. For two donors, a combination of rare alleles could not be resolved, and their DPB1 typings were reported using p-groups.
Conclusion: The assignment of DPB1 typing and correlation to DPB1 unacceptable antigens (UA) in UNet continues to be complex. As evidenced by the absence of some alleles from the Antigen Equivalences table, the identification and utilization of Unacceptable Epitopes is more inclusive, although similarly complex. The option to use p-groups for reporting simplifies the resolution of rare ambiguities and will be particularly useful as the field moves toward high-resolution deceased donor typing.
Methods: HLA typing was performed using LinkSeq HLA Typing Kits (One Lambda) as the primary method. To make rare (non-CWD 2.0) allele assignments, some specimens were additionally tested using Micro SSP (One Lambda), LABType SSO (One Lambda), and/or Olerup SSP (CareDx). 5,542 specimens tested between 01/2023 and 04/2025 at four Eurofins DPT laboratories (Honolulu (2,714), Los Angeles (1,218), Denver (502), and Philadelphia (1,108)) were included in the analysis (numbers in parentheses represent the volumes tested at each lab). Due to the lower numbers of deceased donors in Honolulu, living donor and transplant candidates were also included.
Results: Over 5,000 HLA typings were reviewed. A total of 54 DPB1 alleles were identified (Figure 1). The five most frequently identified alleles represented >75% of all alleles detected. Using CWD 2.0, the frequency catalog utilized in the LinkSeq software, SureTyper, 30 Common, 6 Well-Documented (WD), and 18 non-CWD alleles were detected. However, using the CIWD 3.0.0 catalog, this became 48 Common, 1 Intermediate, 1 WD, and 4 non-CIWD alleles. Each of the non-Common (CIWD 3.0.0) alleles was identified only once. Of the 54 identified alleles, 8 were not included in OPTN Table 4-15: HLA DPB1 Unacceptable Antigen Equivalences. However, all but two of these were only seen once, representing <0.1% of typed alleles. For two donors, a combination of rare alleles could not be resolved, and their DPB1 typings were reported using p-groups.
Conclusion: The assignment of DPB1 typing and correlation to DPB1 unacceptable antigens (UA) in UNet continues to be complex. As evidenced by the absence of some alleles from the Antigen Equivalences table, the identification and utilization of Unacceptable Epitopes is more inclusive, although similarly complex. The option to use p-groups for reporting simplifies the resolution of rare ambiguities and will be particularly useful as the field moves toward high-resolution deceased donor typing.
