(602) A Singleplex HLA Typing Approach for Nanopore Sequencing

Maarten Penning, PhD
– General Manager, GenDx
Speaker(s)
Aim: Long read sequencing technologies are increasingly employed by HLA laboratories. In the field of transplantation diagnostics, multiplex assays are most prevalent in analyzing multiple HLA loci at high efficiency. However, specific HLA loci are increasingly associated with various diseases, resulting in an increased interest in (single locus) HLA typing for disease association testing and companion diagnostics (CDx) purposes. To facilitate this, we are developing whole gene singleplex HLA typing assays for nanopore sequencing, consisting of 10different assays to amplify 11 HLA genes. The combination of single gene amplification and nanopore sequencing, allows for flexible and fast typing results. Amplicons generated with the singleplex assays can be combined with the NGS-Pronto® workflow, introducing flexibility in sequencing of samples and loci.
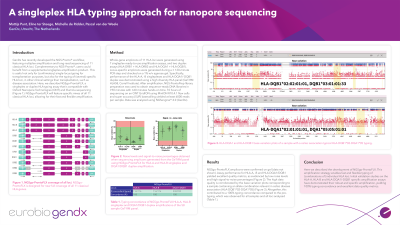
Methods: To determine the performance of the singleplex amplification strategies, DNA from the Genetic Testing Reference Material Coordination Program (GeT-RM) HLA58 panel was used. The 10 different amplification strategies were used to generate whole gene amplicons of the 11 HLA loci, with DRB1 and DRB3 being combined in a single assay due to high homology between these loci. After amplification, amplicons were loaded on a 1% w/v agarose gel to confirm successful amplification and concentrations were determined using Qubit. To limit the number of workflows to be performed, amplicons were pooled in an equimolar ratio per sample followed by library preparation using NGS-ProntoPrep®. Sequencing was performed on a GridION sequencing device using MinION R10.4.1 flow cells andsuper accuracy basecalling. Sequencing for 10 hours (~10 minutes per samples) generated at least 6000 reads per sample. Subsequently, data was analyzed with NGSengine® HLA typing software (GenDx).
Results: Strong bands on gel were observed with correct sizing for each of the 10 assays. NGS sequencing on ONT with subsequent analysis in NGSengine® resulted in a high concordance with available pre-types atthree-field resolution. Typing results were obtained in <5 minutes analysis time per sample for all 58 samples.
Conclusion: In summary, we developed a HLA typing assay designed to be flexible and fast, allowing for typing of single HLA genes at three field resolution.
Methods: To determine the performance of the singleplex amplification strategies, DNA from the Genetic Testing Reference Material Coordination Program (GeT-RM) HLA58 panel was used. The 10 different amplification strategies were used to generate whole gene amplicons of the 11 HLA loci, with DRB1 and DRB3 being combined in a single assay due to high homology between these loci. After amplification, amplicons were loaded on a 1% w/v agarose gel to confirm successful amplification and concentrations were determined using Qubit. To limit the number of workflows to be performed, amplicons were pooled in an equimolar ratio per sample followed by library preparation using NGS-ProntoPrep®. Sequencing was performed on a GridION sequencing device using MinION R10.4.1 flow cells andsuper accuracy basecalling. Sequencing for 10 hours (~10 minutes per samples) generated at least 6000 reads per sample. Subsequently, data was analyzed with NGSengine® HLA typing software (GenDx).
Results: Strong bands on gel were observed with correct sizing for each of the 10 assays. NGS sequencing on ONT with subsequent analysis in NGSengine® resulted in a high concordance with available pre-types atthree-field resolution. Typing results were obtained in <5 minutes analysis time per sample for all 58 samples.
Conclusion: In summary, we developed a HLA typing assay designed to be flexible and fast, allowing for typing of single HLA genes at three field resolution.
